 Data Science
Data Science
Pandas: Advanced booleans
This article is part of a series of practical guides for using the Python data processing library pandas. To see view all the available parts, click here.
In other sections in this series, we’ve looked at how we can use booleans (a value that is either True or False) in pandas. Specifically, we’ve looked at how a list or array of booleans can be used to filter a DataFrame. In those examples we generated lists of booleans using simple comparisons like “are the values in the fixed acidity column > 12?” However, simple comparisons like this are only one of many ways we can create booleans. In this guide we are going to look at a range of methods that allow us to do more complex comparisons, while also making our code more concise and easier to understand.
Data Types
The ways we can generate these booleans varies based on the data type of the column. For example, if I have some text (i.e. a “string”), I might want to check if:
- the string partially matches another string (e.g. does this string contain a certain word?)
- the xth character in the string is a specific character, or
- the string is upper or lower case.
These kinds of checks don’t make sense with a numerical value. Conversely, checking whether one string is greater than another string, although possible, doesn’t really make sense. Below we will walk through some common ways we can form booleans using three different data types, numeric, strings and dates.
Reading in some data
If you don’t have a dataset you want to play around with, University of California Irvine has an excellent online repository of datasets that you can play with. For this explainer we are going to be using the Wine Quality dataset. If you want to follow along, you can import the dataset as follows:
import pandas as pd
df = pd.read_csv("https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv", sep=';')Numerical Data
Starting with numerical data (we are including both floats and integers here), let’s look at some of the comparisons we can make:
Basics
x > y:Trueifxis greater thany.x >= y:Trueifxis greater than or equal toy.x < y:Trueifxis less thany.x <= y:Trueifxis less than or equal toy.x == y:Trueifxis equal toy.x != y:Trueifxis not equal toy.
These basic comparisons work in two different ways. The first is the simple one: comparing the values in a DataFrame column to a single value (df["fixed acidity"] > 12). The second way is comparing the values of two different columns. In this case, the comparison will be done on a row-by-row basis (e.g. df[df["volatile acidity"] < df["citric acid"]]).
Advanced
For numeric data, there are some more advanced comparisons we can make. Often these provide a convenient shorthand for combinations of the basic comparisons shown above.
x.isin([y, z, a]): ReturnsTrueifxexists in a specified list of values. Can be used instead of:x == y | x == z | x == a.
x.between(left=y, right=z, inclusive=True): ReturnsTrueifxis between the values ofyandz(inclusive by default). Can be used instead of:x >= y & x <= zy <= x <= z
x.isna(): ReturnsTrueif x isnull,None,NaN, etc.
Examples
df[df["quality"].isin([3, 8])]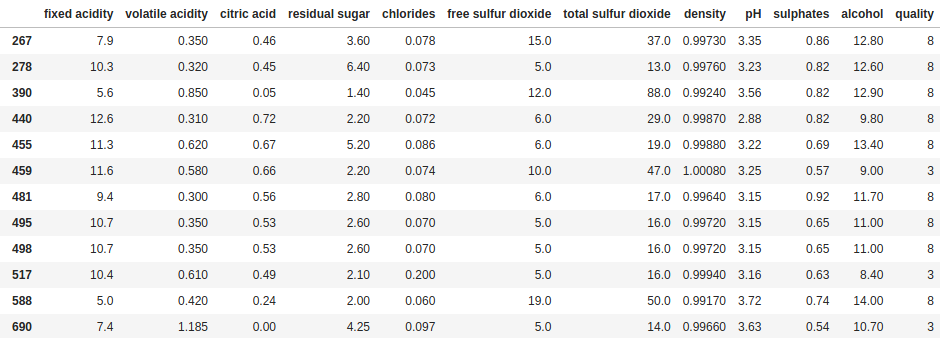
df[df["fixed acidity"].between(11.0, 11.4)]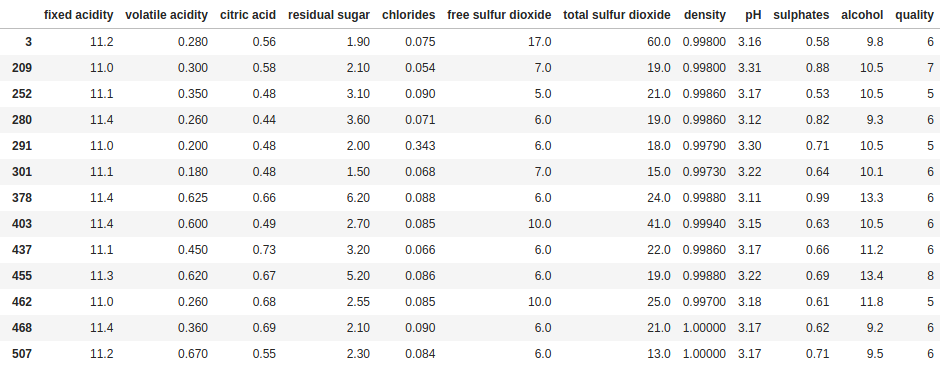
Strings
String comparisons sometimes end up being the most complex that we have to make. Having some basic tools to work with can help to navigate that complexity. But first, let’s create a column with some string data to work with. The following code will create a new column in our DataFrame called random strings and fill it with 20 randomly selected lowercase letters:
import random
import string
df["random strings"] = ["".join(random.choices(string.ascii_lowercase, k=20)) for i in range(len(df))]Basic
x == y:Trueifxis equal toy.x != y:Trueifxis not equal toy.
Technically, you can also compare strings using the < and > operators. This will compare the ASCII value of the characters one by one. However, this is not a common scenario, and if you are going down this path you may be overlooking a cleaner, more robust way to do the comparison.
Advanced
The first thing to note about string methods in pandas is they almost always start with .str. There are many more string specific methods, but the ones we will focus on here are those that we might expect to use in a boolean expression:
df["random strings"].str.contains("abc"): ReturnsTrueif the string contains the pattern"abc".df["random strings"].str.count("abc") > i: ReturnsTrueif the string contains the pattern"abc"more thanitimes.df["random strings"].str.find("abc") > i: ReturnsTrueif the pattern"abc"is found after the ith character in the string. Note,str.findreturns -1 if there is no match.df["random strings"].str.isnumeric() > i: ReturnsTrueif all the characters in the string are numbers. Useful in cases where numeric data might incorrectly be read in as a string or if you have mixed data types.df["random strings"].str.isupper()/.islower(): ReturnsTrueif all the characters in the string are upper case / lower case.
Examples
df[df["random strings"].str.contains("gg")]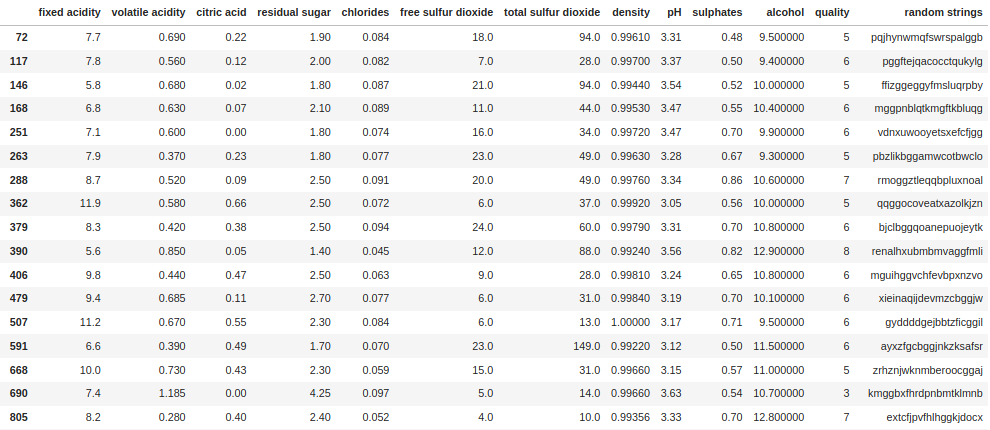
df[df["random strings"].str.count("gg") > 1]
df[df["random strings"].str.find("gg") > 10]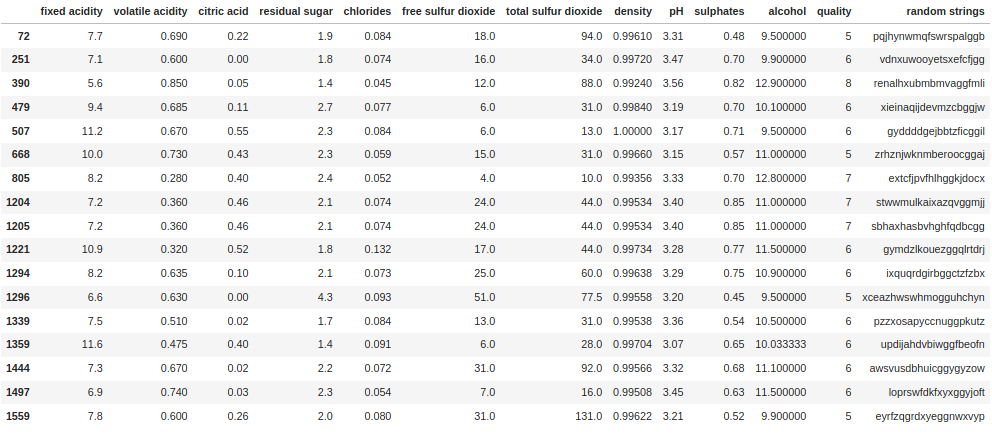
A note on Regex
Several useful str methods such as .str.extract() and .str.match() make use of regular expressions (“regex”), a language used to interact with strings. Regex allows for much more complex string matching based on abstract patterns and character sets, and for the extraction of specific sections of a string. If you are having problems extracting a section of a string, regex is probably what you need. Regex is far too deep a topic to go into here, but there are a range of tools available online which are useful not only for verifying your expressions, but also for learning the basics. One I use regularly is the ExtendsClass Regex Tester.
Dates and Datetimes
For most comparisons, dates and datetimes (dates with a timestamp) work much the same as numeric data. However, there are a couple of important things to be careful of when working with dates and datetimes.
The first is that python has default date and datetime formats, called “date” and “datetime” respectively, both of which are imported from the datetime library (don’t worry, we’ll give an example of this below). When using datetime values in a pandas Series or DataFrame, it will almost always be converted to the pandas equivalent, a timestamp. For practical purposes, you should consider a datetime and a timestamp equivalent.
The second thing to be careful of is comparing dates and timestamps . If you are comparing dates and timestamps stored in the columns of a DataFrame, pandas does a good job of doing the comparisons as you would expect – dropping the timestamp and comparing them on the basis of the date only. However, if you compare a column of timestamps to a single date value, or a column of dates to a single timestamp value, you will get a type error.
Again to get started, let’s create columns with dates and timestamp data to work with. The following code will create two new columns in the DataFrame called random dates and random datetimes and fill them with randomly selected dates:
from datetime import datetime, date
import random
date_list = list(pd.date_range('2017-01-01 23:34:00', periods=len(df)))
random.shuffle(date_list)
df["random datetimes"] = date_list
df["random dates"] = df["random datetimes"].dt.dateWe can check the column data types to make sure they are different:
df[["random datetimes", "random dates"]].dtypes
random datetimes datetime64[ns]
random dates object
dtype: objectWe can also check the values stored in a cell from each column:
df.at[0, "random datetimes"]
Timestamp('2018-04-29 23:34:00')
df.at[0, "random dates"]
datetime.date(2018, 4, 29)There is one very important thing to note here – all the advanced methods we will look at will only work with datetime64 type columns. For a column of dates like we created earlier, there is an easy way to convert them all to timestamps (with the time set to 00:00:00):
df["random dates"] = pd.to_datetime(df["random dates"])Now if we recheck the column data type and data type of the value in the cell:
df[["random datetimes", "random dates"]].dtypes
random datetimes datetime64[ns]
random dates datetime64[ns]
dtype: object
df.at[0, "random dates"]
Timestamp('2018-04-29 00:00:00')Basic
x > y:Trueifxis greater thany.x >= y:Trueifxis greater than or equal toy.x < y:Trueifxis less thany.x <= y:Trueifxis less than or equal toy.x == y:Trueifxis equal toy.x != y:Trueifxis not equal toy.
To reiterate, these comparisons work as they do for numeric values. If the values being compared are both timestamps, the comparison will include the time component. If at least one of the values being compared is a date, the time component will be ignored for the comparison (e.g. "2020-12-01 11:34:00" = "2020-12-01").
Advanced
In addition to the basic comparisons, there are a range of more advanced date specific comparisons we can make. These methods are typically prefaced with .dt and focus on the extraction or manipulation of different parts of a timestamp. Note that the advanced methods we looked at in the numeric section will also work for timestamps.
x.dt.year/quarter/month/day/weekday == y:ReturnsTrueif the extracted date part of the date is equal to i. Works for both dates and datetimesx.dt.hour/minute/second/microsecond == y:ReturnsTrueif the extracted time part of the datetime is equal to i. Only works for datetimes.x.dt.is_month_start/is_month_end:ReturnsTrueif the date is the first/last day of the month. The month end method is often particularly handy.
Examples
df[df["random dates"].dt.year == 2021]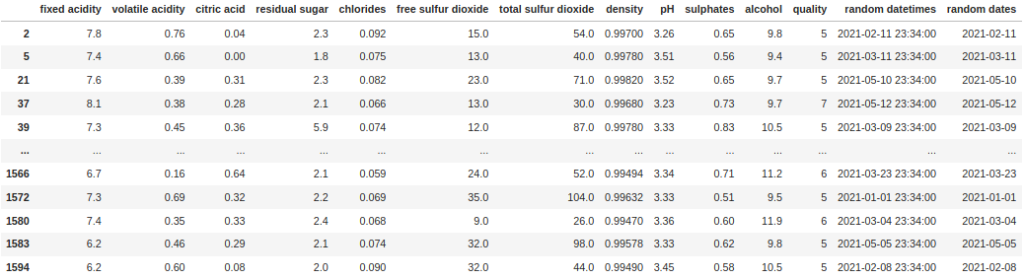
df[df["random datetimes"].dt.is_month_end]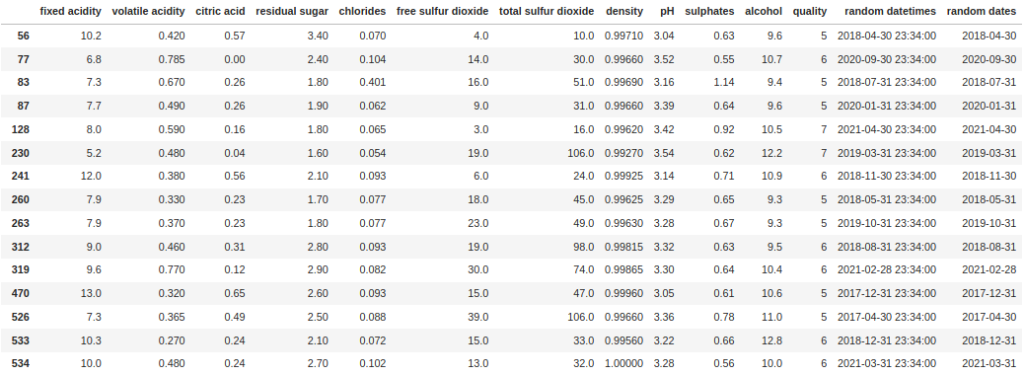
df[df["random datetimes"].between('2020-01-01', '2020-01-10')]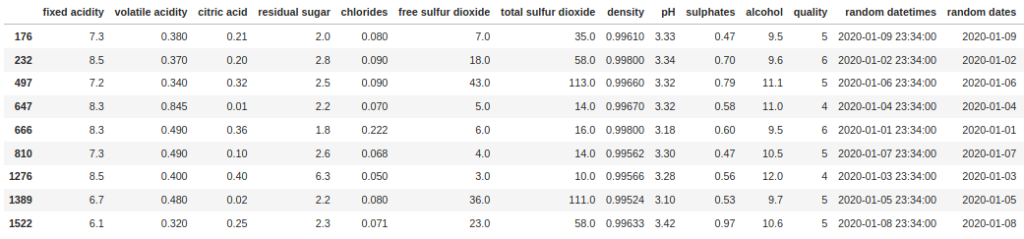
Boolean
What is the difference between a column containing True and False values and the lists of True and False values we use to filter DataFrames? Absolutely nothing! You can use these columns to filter a DataFrame just by passing them as the filter, or if you need to invert the selections (i.e. keep the rows that are False), you can use negation (~). First let’s create a column with random True and False values called random booleans:
df["random booleans"] = [random.random() > 0.5 for i in range(len(df))]Now we can filter on this column just by passing the column as a filter:
df[df["random booleans"]]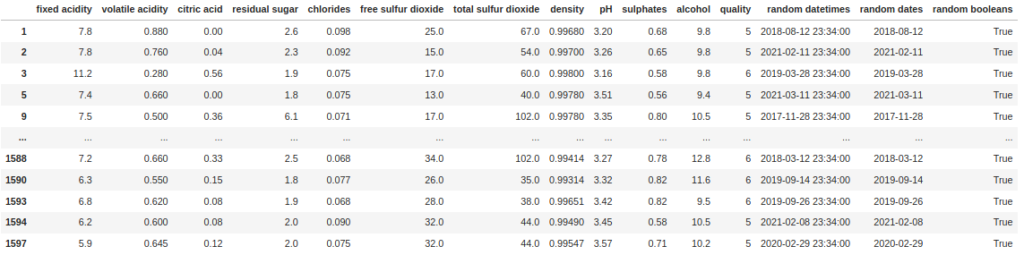
Note that we do not need to do something like df[df["random booleans"] == True]. Likewise if we want to filter for the False rows, we can just use negation (~):
df[~df["random booleans"]]
Wrapping Up
Filtering and segmenting DataFrames is one of the most common tasks when it comes to working with data in pandas. Having a range of methods to apply filters then becomes vital to be an effective data scientist. In this guide we looked at a selection of useful ways to generate filters, covering four different data types (numeric, strings, timestamps and boolean). In addition, we also touched on some of the complexity that comes with these data types (regex with strings, comparability of dates, datetimes and timestamps).
Of course, having access to all these methods from memory alone is something that only comes with significant experience, so initially the going will feel slow. But while you are building up that experience, bookmark this page (or any other reference page you find useful) for quick reference, and soon enough you will find you don’t need to refer back to it.